_Pseudomonas as an opportunistic pathogen
Pseudomonas aeruginosa is an opportunistic pathogen and the leading cause of Gram-negative nosocomial infections, including ventilator-associated pneumonia, infections related to mechanical heart valves, catheters, stents, grafts, sutures, and contact lens-associated corneal infections. Antibiotic therapy can remove planktonic bacteria, but bacteria within biofilms can survive antibiotic treatment. When antibiotic treatment ends, the biofilm can again shed planktonic cells resulting in recurrent acute infection. This cycle of infection is difficult to stop and often requires the removal of the contaminated surface to remove the bacterial biofilm. As with other nosocomial pathogens, the primary reason for the success of P. aeruginosa as an opportunistic pathogen is its ability to form biofilms that provide the bacterium with protection from therapeutic intervention. Biofilm formation by P. aeruginosa is mediated by the production of two extracellular polysaccharides PEL and PSL. In addition, P. aeruginosa is important because it infects more than 90% of cystic fibrosis (CF) patients. Upon mucoid conversion, P. aeruginosa produces copious amounts of alginate, which is associated with decreased lung function and morbidity and mortality of CF patients. Both PEL and alginate polysaccharide synthesis require binding to the secondary signaling molecule cyclic-di-GMP (c-di-GMP). These characteristics make P. aeruginosa one of the best model organisms for the study of c-di-GMP signaling and biofilm mediated chronic infections.
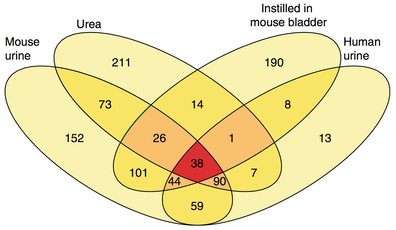 Number of down-regulated genes in P. aeruginosa in response to each condition.
Number of down-regulated genes in P. aeruginosa in response to each condition.
Murine model of catheter-associated urinary tract infection (CAUTI)
Biofilm-based chronic infections are difficult to study due to difficulty of animal models of infection. We have adapted a murine model of CAUTI to study the ability of P. aeruginosa to cause chronic, biofilm infections. We showed that P. aeruginosa forms a biofilm that is independent of known exopolysaccharides required for biofilm formation in laboratory growth media (Cole, et al, Infection and Immunity, 2014). We show that urine and urea, the major nitrogenous waste in urine, can cause P. aeruginosa to switch to a DNA-dependent biofilm from an exopolysaccharide-dependent biofilm. By RNA-seq, we show that P. aeruginosa responds to urine and urea with a set of transcriptomic responses including inhibition of quorum sensing (Figure to the right) and metabolic changes (Cole, Hall, et al, Nature Communications 2018). We are currently studying the effect of these regulatory response during CAUTI.
Biofilm-based chronic infections are difficult to study due to difficulty of animal models of infection. We have adapted a murine model of CAUTI to study the ability of P. aeruginosa to cause chronic, biofilm infections. We showed that P. aeruginosa forms a biofilm that is independent of known exopolysaccharides required for biofilm formation in laboratory growth media (Cole, et al, Infection and Immunity, 2014). We show that urine and urea, the major nitrogenous waste in urine, can cause P. aeruginosa to switch to a DNA-dependent biofilm from an exopolysaccharide-dependent biofilm. By RNA-seq, we show that P. aeruginosa responds to urine and urea with a set of transcriptomic responses including inhibition of quorum sensing (Figure to the right) and metabolic changes (Cole, Hall, et al, Nature Communications 2018). We are currently studying the effect of these regulatory response during CAUTI.
Cyclic-di-GMP regulation and Protein-ligand interactions
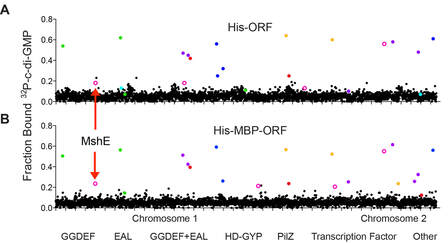
Roelofs, et. al. PLoS Pathogens, 2015
Cyclic-di-GMP is a central signaling molecule that reciprocally regulates the Biological metabolites regulate fundamental biological processes by altering the functions of binding protein partners. The identification of protein-ligand interactions is restricted by the ability of current methodology to detect such measurements in a high-throughput manner. This problem is illustrated by studies of c-di-GMP regulation of bacterial biofilm and virulence. Searches for c-di-GMP binding proteins have yielded one class of PilZ binding proteins that can now be identified through bioinformatics approaches by applying hidden Markov models. However, PilZ domain-containing proteins do not account for all of the functions of c-di-GMP in bacteria. Four other types of c-di-GMP binding proteins, including PelD, FleQ, Clp and VpsT, have been individually discovered by phenotypic or structural studies. Each of these proteins has few amino acids that are required for cdiGMP binding and these amino acid sequences have not been useful for identifying other putative binding proteins through bioinformatic approaches. Thus, neither phenotypic-based studies nor bioinformatic-based approaches have allowed for the systematic identification of all binding proteins of cdiGMP. Similarly, no current methodology exists to comprehensively identify all protein receptors for any other regulatory pathway mediated by a low molecular weight ligand. The identification of additional c-di-GMP binding proteins will reveal the molecular basis of c-di-GMP control of various biological pathways. We have shown that DRaCALA specifically detects c-di-GMP-protein interactions in whole-cell lysates. The ability to detect ligand-protein interactions in whole cells bypasses the requirement to purify proteins and will allow for the screening of open reading frame libraries from various organisms. From a screen for c-di-GMP binding proteins in V. cholerae, we have identified MshE, the ATPase for type IV pili and type II secretion, which the founding member of a new family of c-di-GMP binding domain. We are currently conducting these studies to systematically identify binding proteins for c-di-GMP and other signaling nucleotides.
Differential Radial Capillary Action of Ligand Assay (DRaCALA)
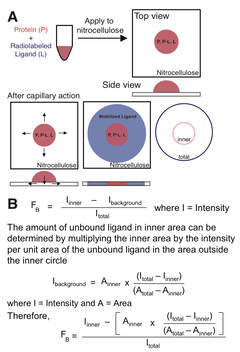
Fig. 1
DRaCALA is a simple, rapid and quantitative technique that allows detection of protein binding to low molecule weight ligands. DRaCALA exploits the ability of nitrocellulose membranes to preferentially sequester proteins over small molecule ligands. When a mixture of protein and radiolabeled ligand is spotted onto a dry nitrocellulose membrane, protein and bound ligand are immobilized at the site of contact while free ligand is mobilized by capillary action with the liquid phase (Fig. 1A). DRaCALA is a rapid assay, as the capillary action can be completed in less than 5 seconds. Since DRaCALA does not utilize a wash step, the pattern of ligand migration allows a rapid detection of both the total ligand and the ligand sequestered by proteins. Using four measurements: 1. Total intensity of ligand, 2. Intensity of bound ligand, 3. Total area, and 4. Area of protein, an accurate determination of fraction bound can be rapidly determined.The principle of DRaCALA was illustrated by measuring ligand binding to known nucleotide binding proteins: P. aeruginosa Alg44-PilZ domain binds cyclic-di-GMP (cdiGMP), E. coli CRP binds cAMP, and E. coli NtrB binds ATP. Radiolabeled ligands were incubated with each of the proteins and the mixtures were spotted on nitrocellulose. After spreading by capillary action, membranes were dried and quantitated by phosphorimager. Maltose binding protein (MBP), which does not bind to any of these small molecules, was used as a control.
As expected, each of the radiolabel signals from MBP mixtures was distributed by capillary action (Fig. 2A). CRP specifically bound cAMP, as demonstrated by the sequestration of the signal, but it did not bind cdiGMP or ATP. NtrB bound ATP, but not cdiGMP or cAMP (Fig. 2A). Similarly, Alg44PilZ bound cdiGMP, but not cAMP or ATP (Fig. 2A). These results and those presented in Roelofs et al, PNAS 2011 demonstrate the power of DRaCALA for detecting protein-ligand interactions.
As expected, each of the radiolabel signals from MBP mixtures was distributed by capillary action (Fig. 2A). CRP specifically bound cAMP, as demonstrated by the sequestration of the signal, but it did not bind cdiGMP or ATP. NtrB bound ATP, but not cdiGMP or cAMP (Fig. 2A). Similarly, Alg44PilZ bound cdiGMP, but not cAMP or ATP (Fig. 2A). These results and those presented in Roelofs et al, PNAS 2011 demonstrate the power of DRaCALA for detecting protein-ligand interactions.
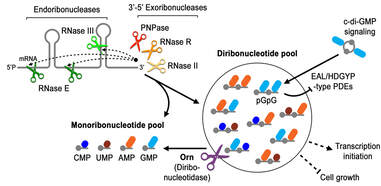 Model for diribonucleotide synthesis, degradation and signaling
Model for diribonucleotide synthesis, degradation and signaling
Degradation of linear diribonucleotides
We have identified oligoribonuclease (Orn) as the primary enzyme that degrades pGpG, the linear degradation product of c-di-GMP (Orr, et al, PNAS 2015). In collaboration with Holger Sondermann (Cornell) and Wade Winkler (UMD), we have revealed that Orn has exceptional specificity for diribonucleotides based on structural, biochemical and microbiological studies (Kim, Lormand, Weiss, et al, eLife 2019). We are currently seeking to understand how diribonucleotides act on the cell.
We have identified oligoribonuclease (Orn) as the primary enzyme that degrades pGpG, the linear degradation product of c-di-GMP (Orr, et al, PNAS 2015). In collaboration with Holger Sondermann (Cornell) and Wade Winkler (UMD), we have revealed that Orn has exceptional specificity for diribonucleotides based on structural, biochemical and microbiological studies (Kim, Lormand, Weiss, et al, eLife 2019). We are currently seeking to understand how diribonucleotides act on the cell.
Small molecule inhibitors of bacteria virulence
We have previously used cell based assays to identify small molecule inhibitor of Type III secretion substrates, ExoU and ExoS. Currently we are interested in using a biochemical approach, such as DRaCALA, to identify small molecular inhibitors.
Publications
Degradation of pGpG and linear dinucleotides
Cyclic-di-GMP and cyclic-di-AMP regulation
Catheter-associated urinary tract infection (CAUTI)
Pseudomonas biology and pathogenesis
Small molecule inhibitors
- Kim, S.K., Lormand, J.D., Weiss, C.A., Eger, K.A., Turdiev, H., Turdiev, A., Winkler, W.C., Sondermann, H., Lee, V.T. 2019. A dedicated diribonucleotidase resolves a key bottleneck as the terminal step of RNA degradation. Elife. 8. pii: e46313. PDF
- Orr, M.W., Weiss, C.A., Severin, G.B., Turdiev, H., Kim, S.K., Turdiev, A., Liu, K., Tu, B.P., Waters, C.M., Winkler, W.C., Lee, V.T. 2018. A Subset of Exoribonucleases Serve as Degradative Enzymes for pGpG in c-di-GMP Signaling. J. Bacteriol. 2018 Nov 26;200(24). pii: e00300-18. PDF
Cyclic-di-GMP and cyclic-di-AMP regulation
- Hall, C.L. and Lee, V.T. 2018. Cyclic-di-GMP regulation ofvirulence in bacterial pathogens. Wiley Interdiscip Rev RNA. 2018 Jan;9(1). doi: 10.1002/wrna.1454. PDF
- Roelofs, K.G., Jones, C.J., Helman, S.R., Shang, X., Orr, M.W., Goodson, J.R., Galperin, M.Y., Yildiz, F.H., and Lee, V.T. 2015. Systematic Identification of Cyclic-di-GMP Binding Proteins in Vibrio cholerae Reveals a Novel Class of Cyclic-di-GMP-Binding ATPases Associated with Type II Secretion Systems. PLoS Pathogens. 11(10):e1005232. PDF
- Orr, M.W., Donaldson, G.P., Severin, G.B., Wang, J., Sintim, H.O., Waters, C.M., and Lee, V.T. 2015. Oligoribonuclease is the primary degradative enzyme for pGpG in Pseudomonas aeruginosa that is required for cyclic-di-GMP turnover. Proc Natl Acad Sci U S A. 112(36):E5048-57. PDF
- Corrigan, R.M., Campeotto, I., Jeganathan, T., Roelofs, K.G., Lee, V.T. and Gründling, A. 2013. Systematic identification of conserved bacterial c-di-AMP receptor proteins. Proc Natl Acad Sci U S A. 110(22):9084-9. PDF
- Roelofs, K.G., Wang, J., Sintim, H.O., and Lee, V.T. 2011. Differential radial capillary action of ligand assay for high-throughput detection of protein-metabolite interactions. Proc Natl Acad Sci U S A. 108 (37) 15528-15533. PDF
- Luo, Y., Zhang, J., Watt, S.K., Lee V.T., Sintim H.O. 2011. Differential binding of 2’-biotinylated analogs of c-di-GMP with c-di-GMP riboswitches and binding proteins. Moleular BioSystems. DOI: 10.1039/C2MB05338A. Online link
- Wang, J., Zhou, J., Donaldson, G., Nakayama, S., Yan, L., Lam, Y-F., Lee V.T., Sintim, H.O. 2011. Conservative change to the phosphate moiety of c-di-GMP remarkably affects its polymorphism and binding abilities to DGC, PDE and PilZ proteins. J Am Chem Soc. 133(24):9320-30. Online link
- Lee, V.T., Matewish, J.M., Kessler, J.L. Hyodo, M., Hayakawa, Y., and Lory, S. 2007. A Cyclic-di-GMP Receptor Required for Bacterial Exopolysaccharide Production. Mol. Microbiol. 65: 1474-1484. PDF
- Merighi, M., Lee, V.T., Hyodo, M., Hayakawa, Y., and Lory, S. 2007. The second messenger bis-(3’-5’)-cyclic-GMP and its PilZ domain-containing receptor Alg44 are required for alginate biosynthesis in Pseudomonas aeruginosa. Mol. Microbiol. 65: 876–895. PDF
- Kulasakara, H., Lee, V., Brencic, A., Liberati, N., Urbach, J., Miyata, S., Lee, D.G., Neely, A.N., Hyodo, M., Hayakawa, Y., Ausubel, F.M., and Lory, S. 2006. Analysis of Pseudomonas aeruginosa diguanylate cyclases and phosphodiesterases reveals a role for bis-(3'-5')-cyclic-GMP in virulence. Proc Natl Acad Sci U S A. 103:2839-44. PDF
Catheter-associated urinary tract infection (CAUTI)
- Cole, S.J., Hall, C.L., Schniederberend, M., Farrow III, J.M., Goodson, J.R., Pesci, E.C., Kazmierczak, B.I., Lee V.T. 2018. Host suppression of quorum sensing during catheter associated urinary tract infections. Nature Comm. 9(1):4436. PDF
- Cole, S.J., Lee, V.T. 2015. Cyclic-di-GMP signaling contributes to Pseudomonas aeruginosa mediated catheter-associated urinary tract infection. J Bacteriol. JB.00410-15. PDF
- Cole, S.J., Records, A.R., Orr, M.W., Linden, S.B., and Lee, V.T. 2014. Catheter-associated urinary tract infection by Pseudomonas aeruginosa is mediated by exopolysaccharide-independent biofilms. Infect Immun. 82(5):2048-58. PDF
Pseudomonas biology and pathogenesis
- Salzberg, S., Sommer, D.D., Puiu, D., and Lee, V.T. 2008. Gene-boosted assembly of a novel bacterial genome from very short reads. PLoS Computational Biology. 4:e1000186. Online link
- Lee, V.T., Smith, R.S., Tümmler, B., and Lory, S. 2005. Activities of Pseudomonas aeruginosa effectors secreted by the Type III Secretion system in vitro and during infection. Infect. Immun. 73:1695-705. PDF
- Wolfgang, M.C., Lee, V.T., Gilmore, M.E. and Lory, S. 2003. Coordinate Regulation of Bacterial Virulence Genes by a Novel Adenylate Cyclase-Dependent Signaling Pathway. Dev. Cell. 4:253-63. PDF
Small molecule inhibitors
- Zhou, E., Seminara, A.B., Kim, S.K., Hall, C.L., Wang, Y., Lee V.T. 2017. Thiol-benzo-triazolo-quinazolinone Inhibits Alg44 Binding to c-di-GMP and Reduces Alginate Production by Pseudomonas aeruginosa. ACS Chemical Biol. 15;12(12):3076-3085. PDF
- Lieberman, O.J., Orr, M.W., Wang, Y., and Lee, V.T. 2014. High-throughput screening using the differential radial capillary action of ligand assay identifies ebselen as an inhibitor of diguanylate cyclases. ACS Chemical Biology. 9(1):183-92. Online link
- Lieberman, O.J., DeStefano, J.J., and Lee, V.T. 2013. Detection of cyclic diguanylate G-octaplex assembly and interaction with proteins. PLoS One. 8(1):e53689. doi: 10.1371/journal.pone.0053689. Online link
- Donaldson, G.P., Roelofs, K.G., Luo, Y., Sintim, H.O., Lee, V.T. 2011. A rapid assay for affinity and kinetics of molecular interactions with nucleic acids. Nucleic Acids Research. DOI: 10.1093/nar/gkr1299. PDF
- Smith, J.A.I., Wang, J., Nguyen-Mau, S.-M., Lee, V. and Sintim, H. O. 2009. Facile syntheses of AI-2 and analogues facilitate the identification of cyclohexyl AI-2 as a suppressor of AI-2-mediated processes in Vibrio harveyi. Chemical Communications. DOI:10.1039. Online link
- Lee, V.T. and Kessler, J.L 2009. Type 3 secretion systems as targets for novel antibiotics. IDrugs. 2009. 12(10):636-641.
- Wang, J., Lee, V., Sintim, H.O. 2009. Efforts towards the identification of simpler platensimycin analogs - the total synthesis of oxazinidinyl platensimycin. Chem. Eur. J. 15:2747-50. PDF
- Arnoldo, A., Curak, J., Kittanakom, S., Chevelev, I., Lee, V.T., Sahebol-Amri, M., Koscik, B., Ljuma, L., Roy, P.J., Bedalov, A., Nislow, C., Merrill, R.A., Lory, S. and Stagljar, I. 2007. Identification of small molecule inhibitors of Pseudomonas aeruginosa Exotoxin S using a yeast phenotypic screen. PLoS Genetics. 2:e1000005. Online link
- Lee, V.T., Pukatzki, S., Sato, H., Kikawada, E., Kazimirova, A.A., Huang, J., Li, X., Arm, J.P., Frank, D.W., and Lory, S. 2007. Pseudolipasin A Is a Specific Inhibitor for Phospholipase A2 Activity of Pseudomonas aeruginosa Cytotoxin ExoU. Infect. Immun. 75: 1089-98. PDF
Full list of publications
Funding
NIH NIAID/NIDDK and Cystic Fibrosis Foundation

